Loading...


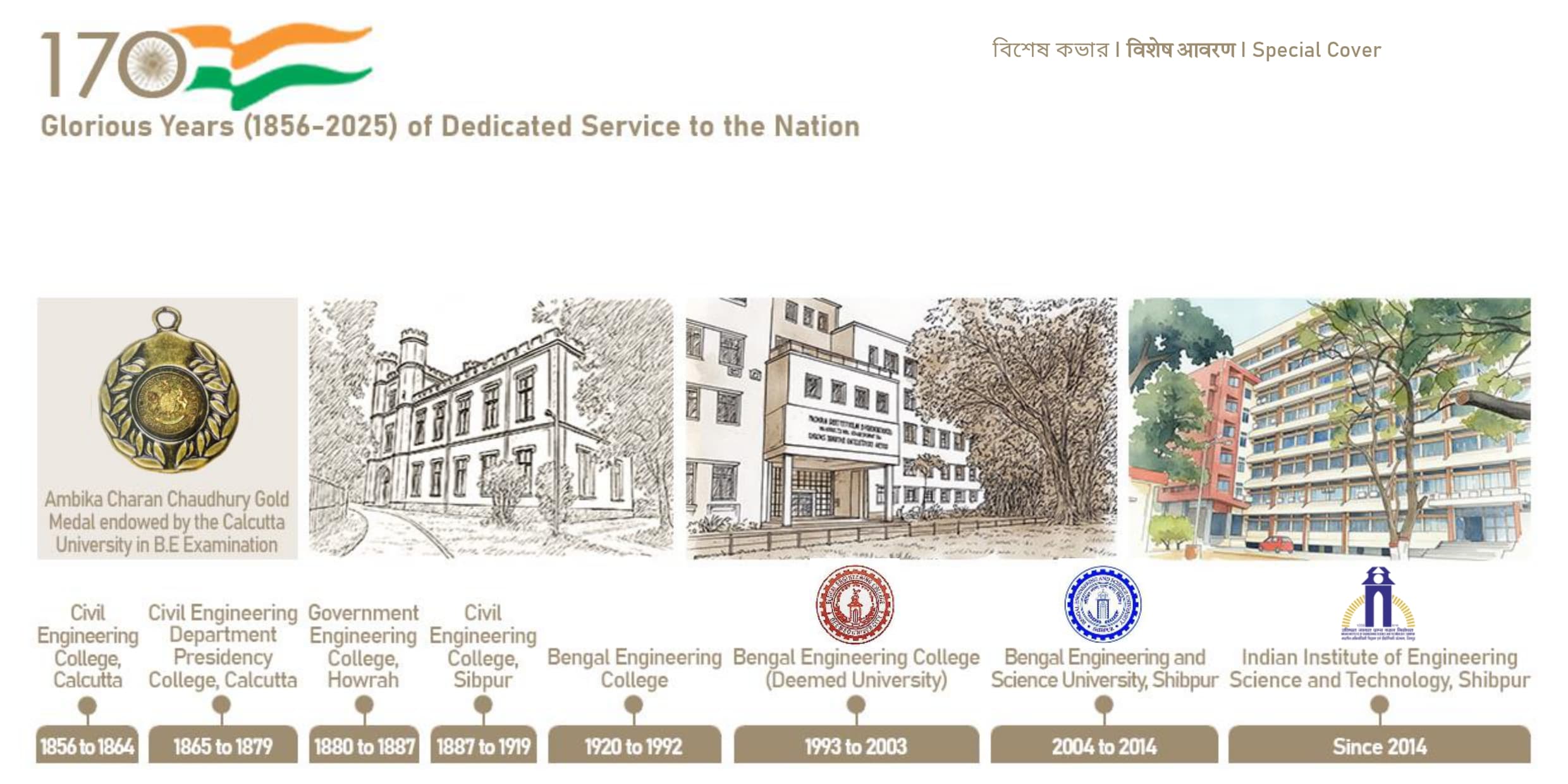
Glorious Years (1856-2025) of Dedicated Service to the Nation


Glorious Years (1856-2025) of Dedicated Service to the Nation
Loading...
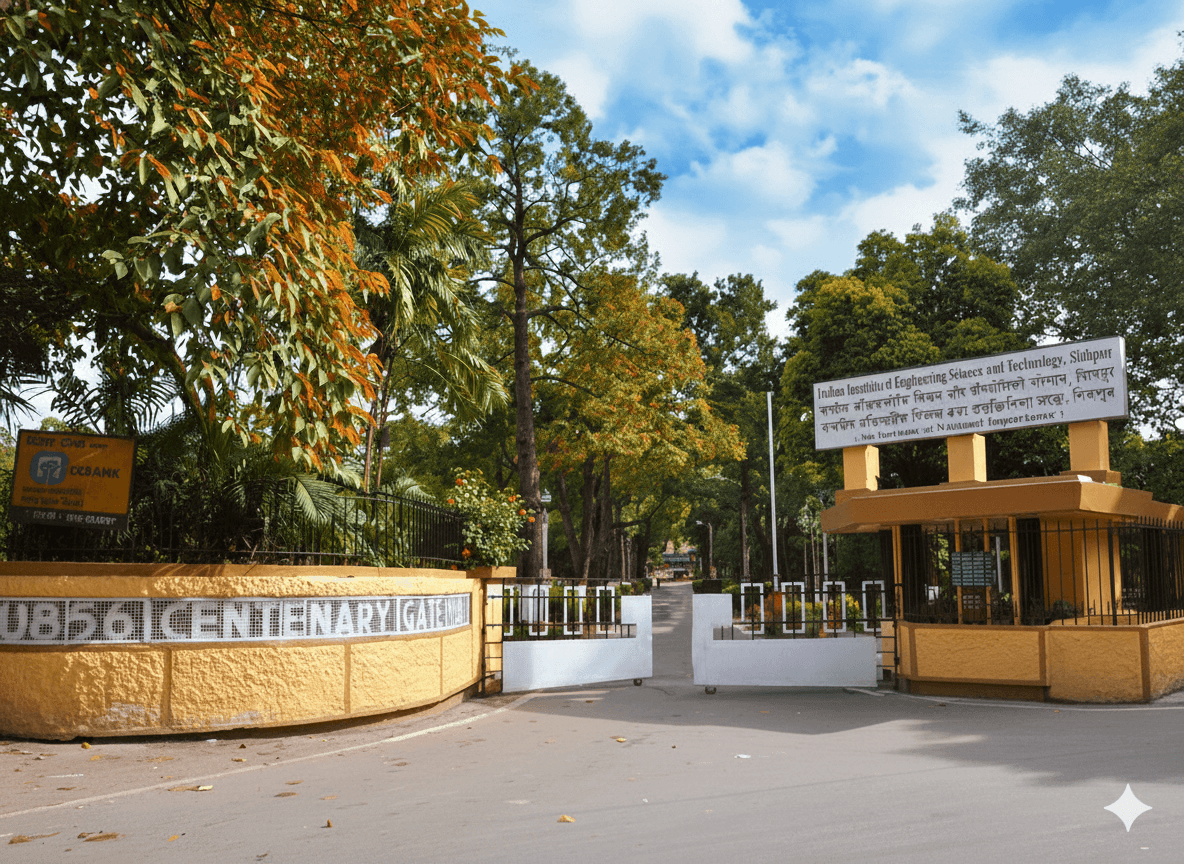
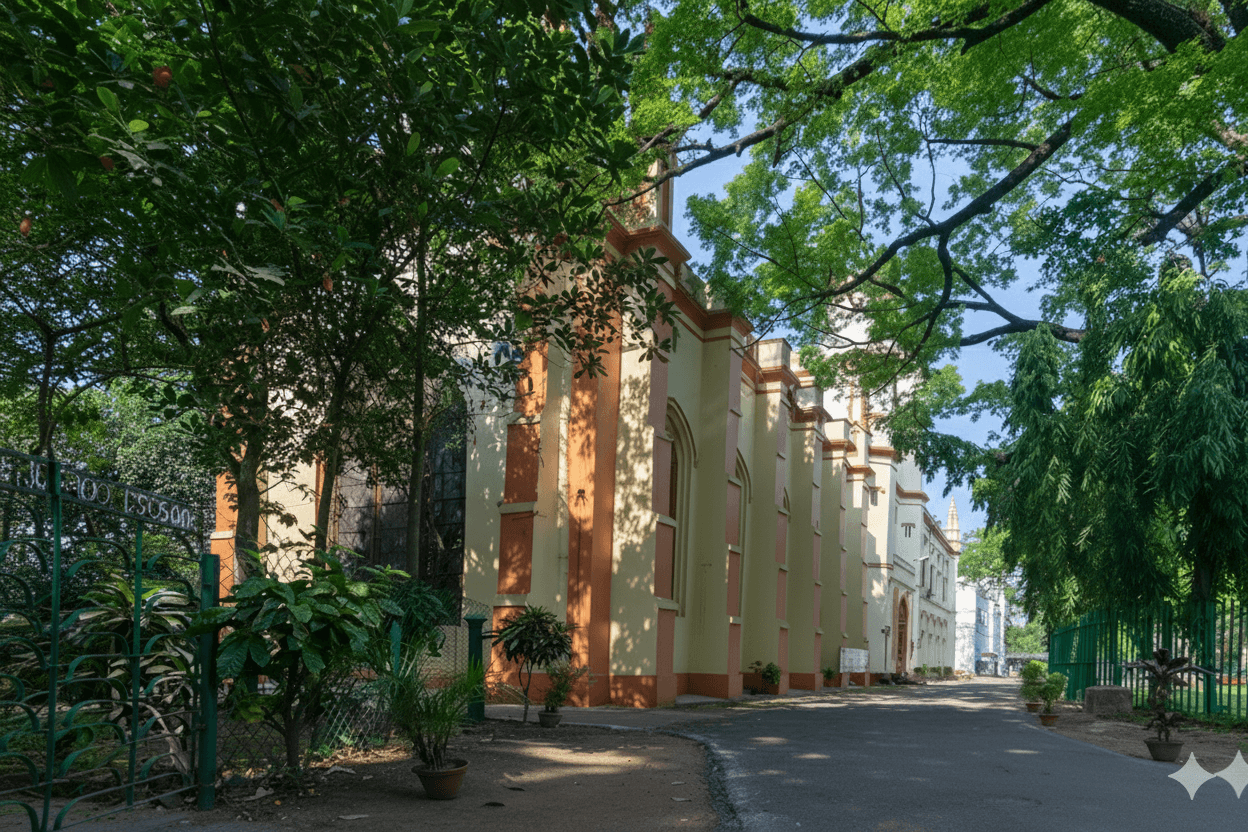


IIEST, Shibpur, is recognized as an Institute of National Importance. Established in 1856, it is India’s second-oldest engineering college. In March 2014, Bengal Engineering and Science University, Shibpur was taken over by the Government of India and converted into an Institute of National Importance through an act of parliament and renamed as Indian Institute of Engineering Science and Technology, Shibpur.It is the first college to become the Indian Institute of Engineering Science and Technology (IIEST) in India in 2014. The Institute has a rich history. It has been empowering the nation since 1856. Considering the year of establishment, it is the third engineering college in India, but considering the year of graduation, it is the second oldest engineering college in India. The Institute has 16 departments and 8 schools, with over 250 faculty members and more than 4000 students.
A rich tradition of art, festivals, and community.

Experience the vibrant campus life and heritage.
4,264
Students Enrolled
827
Research Scholars
245
Faculty Members
238
Non Faculty Members
669
Awards Won
8
Degree Programs

Featured discoveries and innovations from IIEST Shibpur